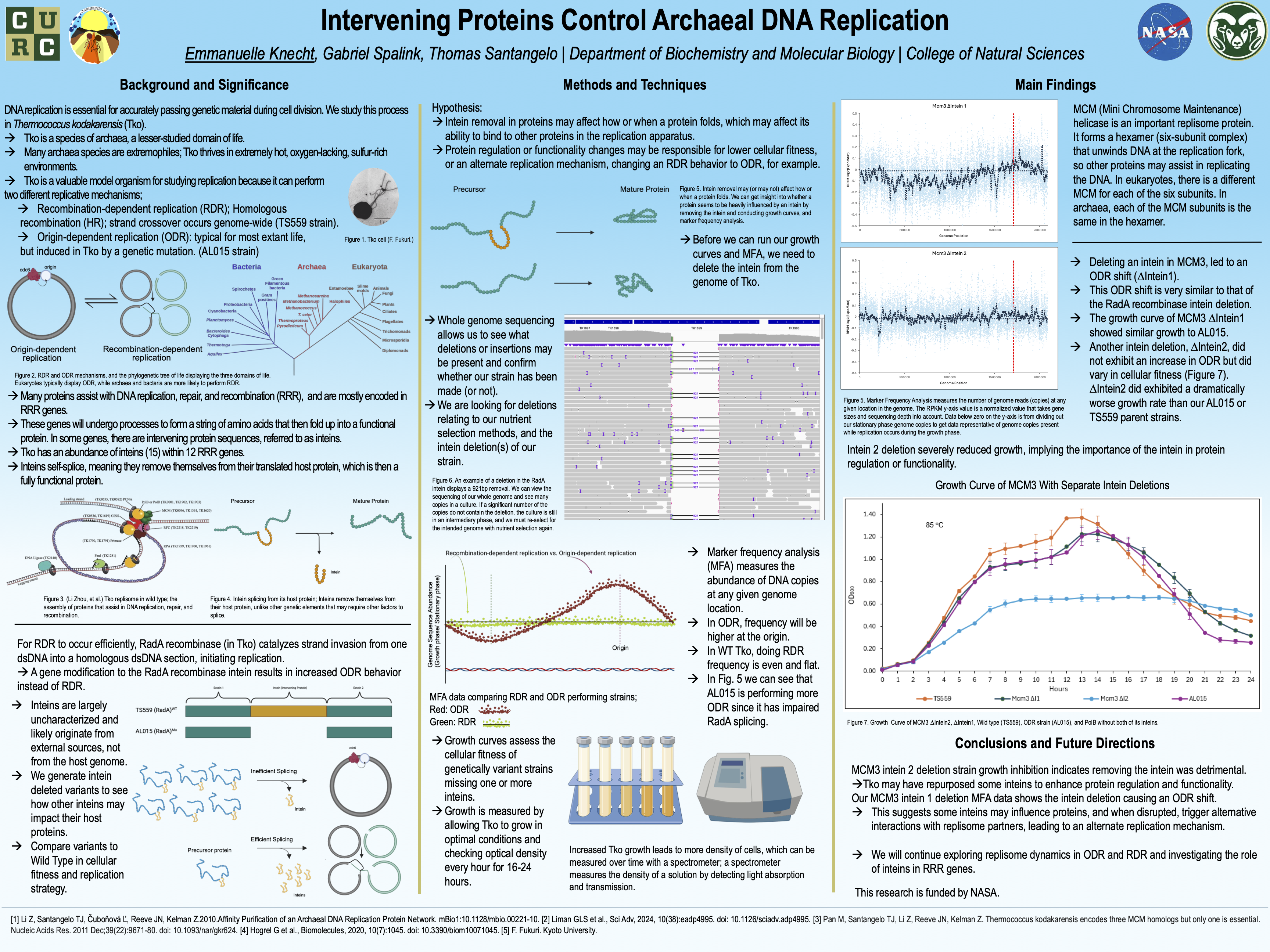
Intervening Proteins Control Archaeal DNA Replication
Category: Research Poster
Author(s): Emmanuelle Knecht
Presenter(s): Emmanuelle Knecht
Mentors(s): Gabriel Spalink, Thomas Santangelo
DNA replication is fundamental for cellular division and genome integrity, where errors result in disease states, including cancer. Given the importance of DNA replication, a variety of proteins assemble in a necessary replication apparatus we call the replisome. The replisome orchestrates DNA replication efficiently and resolves associated errors. The replisome proteins are encoded in replication, repair, and recombination (RRR) genes. Many of these genes also encode special intervening proteins known as inteins, which function by removing themselves from their host protein, and likely originated from unknown external sources rather than the host genome. Our organism of study, Thermococcus kodakarensis (Tko), is an archaeon, in a different domain than eukarya and bacteria, and is adapted to extreme environments. In the genome of Tko, an abundance of inteins are encoded within RRR genes. The function of these inteins remains largely unknown. To better understand how Tko may use inteins, we are constructing genetically modified variants with individual intein deletions to see the effects on replication strategy and cellular fitness. We have already made progress on mapping the Thermococcus kodakarensis replisome, particularly in protein-protein interactions that form the DNA replication apparatus in the preferred replication method of Tko. We are now expanding on our work to elucidate the replisome, with the discovery that Tko can be induced into an alternative replication technique, closer to that of how humans replicate DNA. Our current endeavors in replisome mapping focus on examining protein-protein interactions in both replication behaviors to gain a comprehensive understanding of replisome dynamics.
