Expansion of DNA Sequence-Structure Map by X-Ray Crystallography of Self-Complementary Sequence d(CCCCTAGGGG)
Category: Research Poster
Author(s): Sarah Abercrombie
Presenter(s): Sarah Abercrombie
Mentors(s): Pui Ho
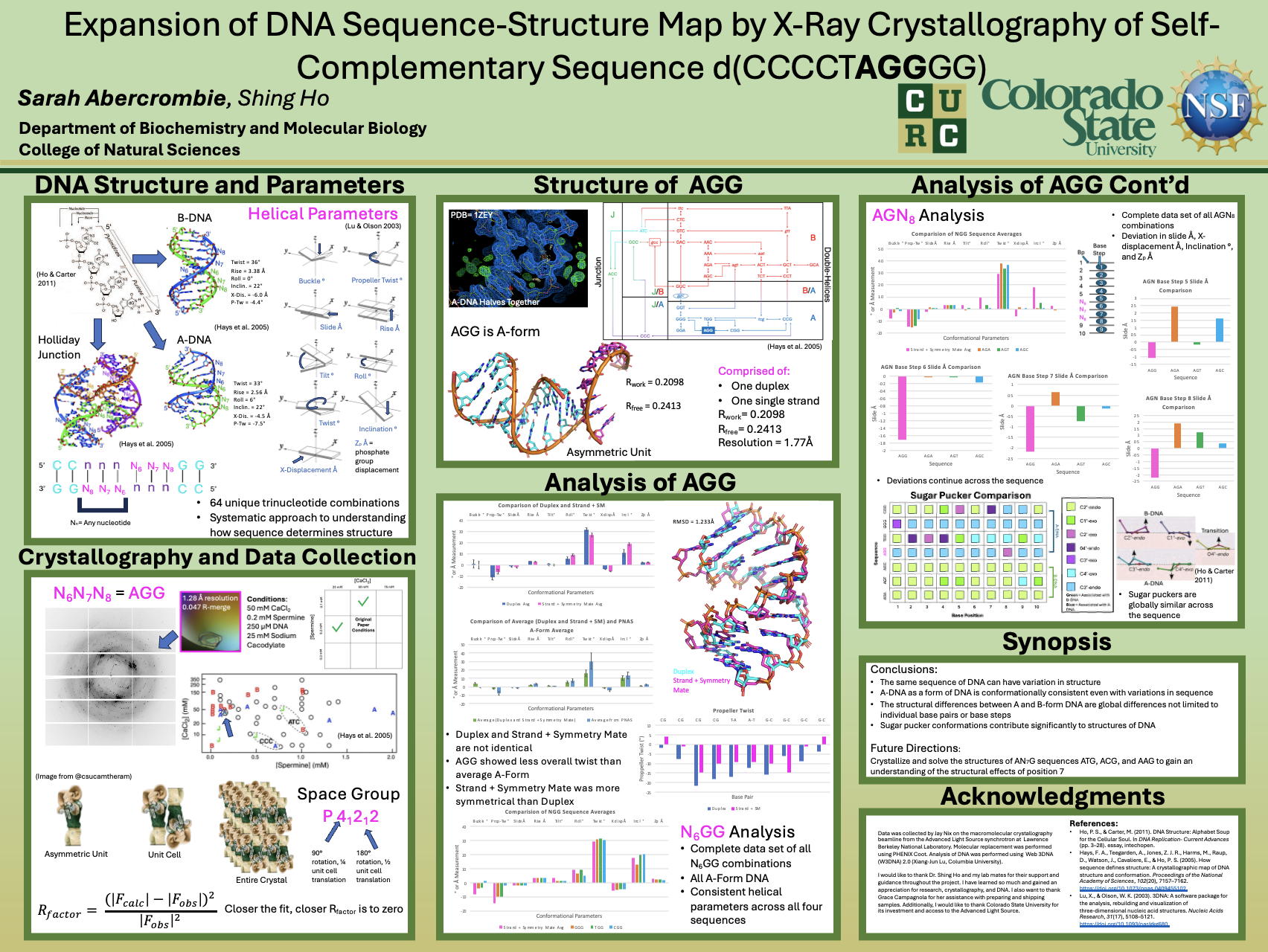
DNA is a polymorphic molecule capable of adopting various forms depending on its sequence. Mapping sequence and form in a systematic manner would be beneficial for understanding how sequence impacts structure and being able to predict specific locations of DNA forms within the genome. The ability to make these predictions would offer insights into how DNA structure regulates processes like replication and transcription. Previous studies have shown the inverted repeat sequence motif d(CCnnnN6N7N8GG), where N6, N7, and N8 can represent any of the four nucleotides, serves as a scaffold for determining the form and structure of all 64 unique possible trinucleotides. One unsolved N6N7N8 trinucleotide is AGG. AGG crystallized under conditions consistent with B-form DNA, resulting in a crystal with a unit cell of 44.64Å x 44.64Å x 76.38Å and a space group of P41212. The structure of the asymmetric unit, solved using molecular replacement, consists of a duplex and a single strand of DNA. Both the duplex and single strand, along with the symmetry mate, are A-form DNA. Comparison of N6GG sequences to AGG showed similar helical parameter values, indicating that A-DNA is overall conformationally consistent. An AGN8 comparison showed AGG had a significant deviation of the amount base steps are slid out, inclined, and positioned away from the center of the helix. This suggests that the G positioned at N8 had global effects on the structure of the helix. This sequence exemplifies how trinucleotide sequence influences structure and the potential structural variation depending on the nucleotide combination.
