Minimum Sequence Requirements of Prion Like Domains in Yeast for Stress Granule Localization
Category: Research Poster
Author(s): Alessandra Donev
Presenter(s): Alessandra Donev
Mentors(s): Eric Ross
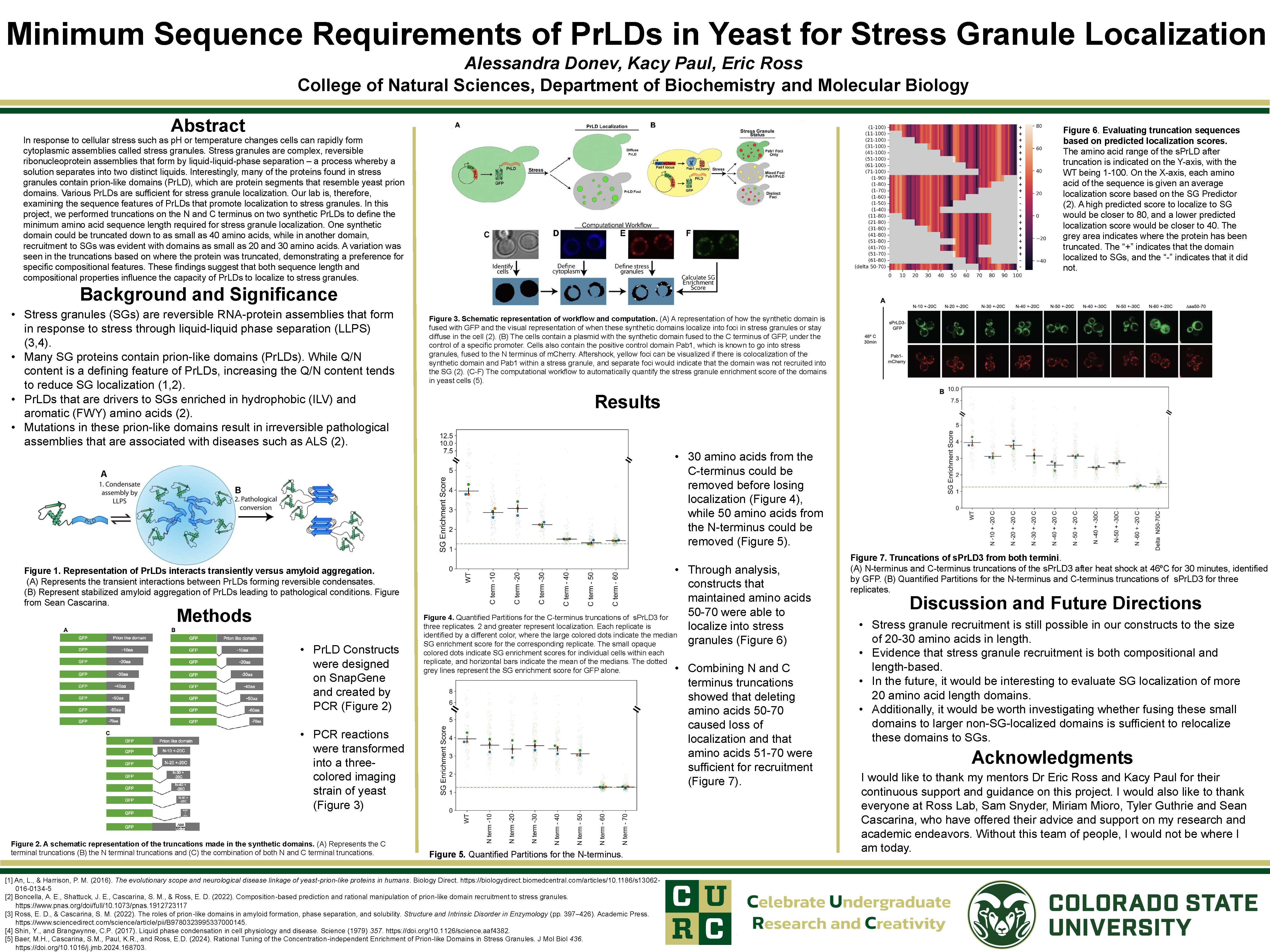
In response to cellular stress such as pH or temperature changes cells can rapidly form cytoplasmic assemblies called stress granules. Stress granules are complex, reversible ribonucleoprotein assemblies that form by liquid-liquid-phase separation – a process whereby a solution separates into two distinct liquids. Interestingly, many of the proteins found in stress granules contain prion-like domains (PrLD), which are protein segments that resemble yeast prion domains. Various PrLDs are sufficient for stress granule localization. Our lab is, therefore, examining the sequence features of PrLDs that promote localization to stress granules. In this project, we performed truncations on the N and C terminus on two synthetic PrLDs to define the minimum amino acid sequence length required for stress granule localization. One synthetic domain could be truncated down to as small as 40 amino acids, while in another domain, recruitment to SGs was evident with domains as small as 20 and 30 amino acids. A variation was seen in the truncations based on where the protein was truncated, demonstrating a preference for specific compositional features. These findings suggest that both sequence length and compositional properties influence the capacity of PrLDs to localize to stress granules.
