Targeting intergenic spaces for genetic improvement of broad-spectrum disease resistance in rice
Category: Research Poster
Author(s): Ellie Misra-Matson
Presenter(s): Ellie Misra-Matson
Mentors(s): Federico Martin
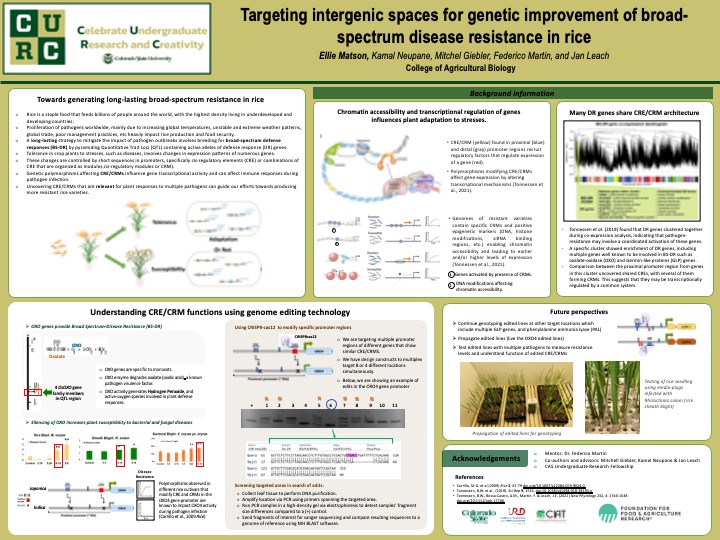
Rising global temperatures and erratic weather patterns drive ecosystem changes allowing plant pathogens to spread and thrive in new niches further stressing agricultural systems. Efforts to contain the spread of plant diseases often rely on the use of agrichemicals. A more sustainable alternative is to enhance natural plant resistance to pathogens and pests. Though, this is a complex and lengthy process that can take up to 10 years. Tolerance in crop plants to stresses involves changes in expression patterns of numerous genes. These changes are controlled by short sequences in promoter areas, specifically cis-regulatory elements (CRE) or assemblies of CRE organized as modules (cis-regulatory modules or CRM). Increasing evidence shows that conserved CRE and CRM are found in promoters of many genes that are co-activated by a single stress, and that they are common to genes co-activated in plants with enhanced tolerance to different stresses. We hypothesize that conserved CRE/CRM enable coordinated gene activation, contributing to enhanced defense responses. Previous work in our group identified multiple genes involved in defense responses including phenylalanine ammonia lyase (PAL), oxalate oxidase (OXO) and germin-like proteins (GLP) which share similar CRMs in the proximal promoter regions. Using CRISPR-cas genome editing technology, we have targeted and modified some of these CRMs in a multiplex approach. Currently, we are genotyping targeted regions to identified beneficial modifications and selecting specific lines for future studies. Edited lines will be tested with different pathogens to analyze their susceptibility response. Our goal is to better understand the role that specific CRMs have in gene activation during defense responses and use them to guide genetic selection of the most actives allele for efficient development of climate-ready varieties.
